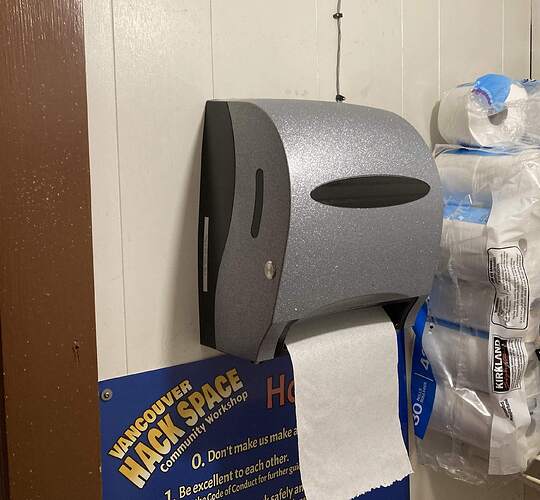
A while back a member (forgive me as I have forgotten who did and can’t find the posts in slack) purchased some paper towel dispensers that looked Cylon’ish…
That got me thinking and the end result is installed in the bathrooms…
Here they are in action…
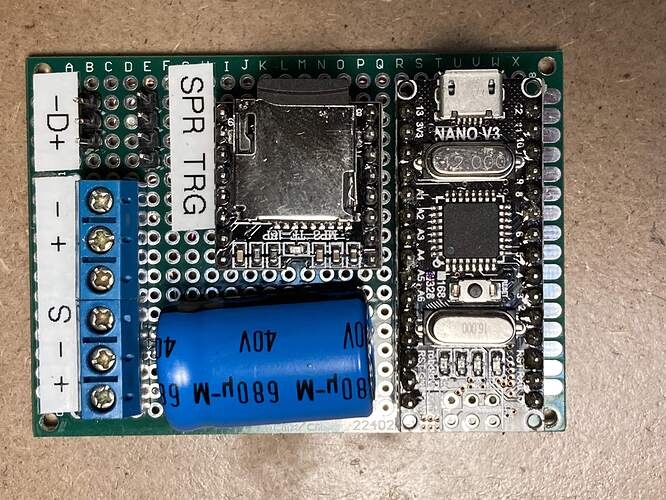
I used a NANO and a DF Player (MP3 player module) and cobbled some code together based on what I could find with Google
VHS_CYLON_6.zip (3.2 KB)
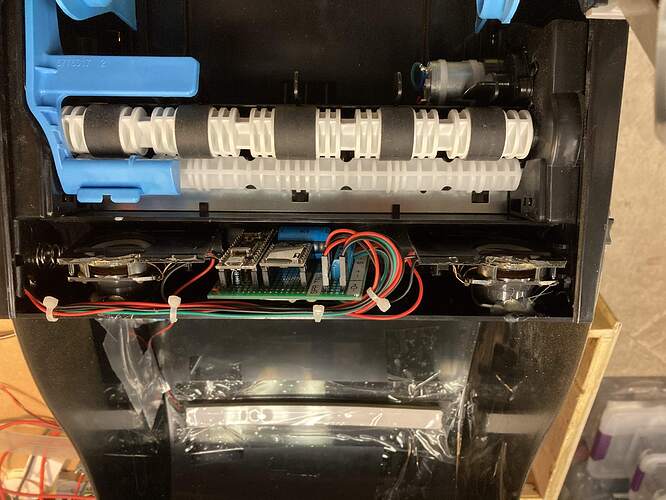
The dispensers have a small motor controlled by a capacitive sensor plate located on the bottom. I put an opto coupler across the motor terminal so the NANO code is triggered when the motor is started. The code then plays the sound (1 of 34 stored on a SD card) and runs the Pixel LEDS in a Larson Scanner effect till the audio is completed.
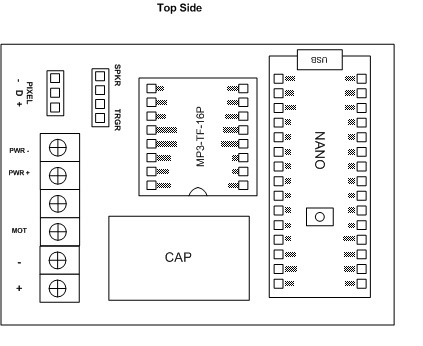
I don’t have a circuit diagram but the code is reasonably documented…
I added a pushbutton trigger on the side so you don’t have to use up paper towel should you want to activate it…
The units normally have battery power but I used the battery holder to house the board and speakers so ended up powering it from a 5V power supply located on the wall above the opening to the bathrooms.
So now we will never have to replace the batteries. The sound volume is a bit muffled (I used some old laptop speakers I found at the Space) but I can crank the volume a bit if needed (it is controlled in the code). The sound bites are just stuff I found that seemed appropriate but can be replaced or added to (PSA mebbe??).
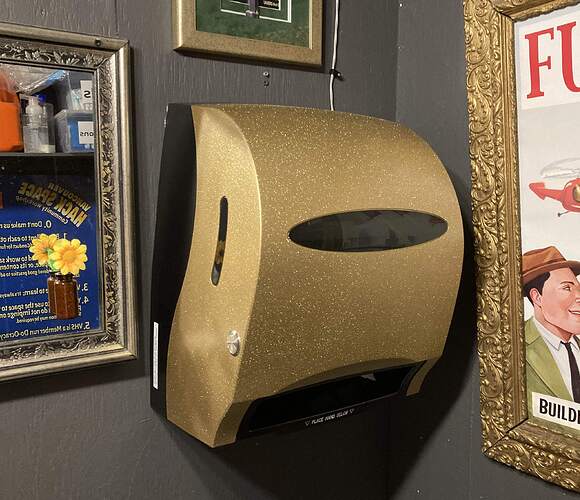
And once I installed the guts I painted them with some metallic sparkle paint I had…




